Nikolai Slavov
COE Distinguished Professor,
Bioengineering
Founding Director,
Parallel Squared Technology Institute
Allen Distinguished Investigator,
Allen Frontiers Group
Director,
Single-Cell Proteomics Center
Affiliated Faculty,
Biology
Affiliated Faculty,
Chemistry and Chemical Biology
Office
- Mugar Life Sciences, Office 334
- 617.373.6405
Research Focus
Single-cell proteomics, immunology, cancer drug resistance, translational regulation, quantitative systems biology, mass spectrometry, RNA decoding, protein synthesis and degradation, macrophage polarization, aging, neurodegenerative diseases
About
Nikolai Slavov received undergraduate education from MIT and a doctoral degree from Princeton University for characterizing the coordination of cellular growth with gene expression and metabolism.
The Slavov laboratory pioneered experimental and computational methods for single-cell proteomics and used them to connect protein covariation across single cells to functional phenotypes, including macrophage polarization, emergence of drug resistance priming, early mammalian development, proteostasis, neurodegenerative diseases, and stem cell differentiation. These technologies provided a foundation for establishing Parallel Squared Technology Institute (PTI).
Prof. Slavov organizes the annual single-cell proteomics conference and contributes to organizing other leading conferences.
Education
- PhD (2010), Botstein Laboratory, Princeton University
- BS (2004), Biology, Massachusetts Institute of Technology
Honors & Awards
- Biemann Medal by the American Society For Mass Spectrometry
- 2026 Distinguished Faculty Award
- HUPO Discovery in Proteomic Sciences Award
- 2022 College of Engineering Faculty Fellow
- Allen Distinguished Investigator Award
- Excellence in Mentoring Award
- NIH Director’s New Innovator Award
- SPARC Award from the Broad Institute of MIT and Harvard
- Princeton University Dean’s Award
- IRCSET Postgraduate Research Fellowship
- Finalist in the Young European Entrepreneur Competition
- Princeton Graduate Fellowship
- MIT Undergraduate Fellowship
- Eureka Fellowship for Academic Excellence
- Bronze Medal in the 31st International Chemistry Olympiad
- National Diploma for Exceptional Achievements in Chemistry
Teaching Interests
Leadership Positions
- Founding director of Parallel Squared Technology Institute
- Established the annual Single-cell Proteomics conference
- Co-organized workshops at scientific conferences, including NeurIPS (Learning Meaningful Representations of Life), and HUPO
Professional Affiliations
- Founding director of Parallel Squared Technology Institute (PTI)
- Founding director of Single-Cell Proteomics Center
- Barnett Institute of Chemical and Biological Analysis
- Broad Institute of MIT & Harvard
- American Society for Mass Spectrometry (ASMS)
- American Society for Cell Biology (ASCB)
- Genetics Society of America (GSA)
Research Overview
Single-cell proteomics, immunology, cancer drug resistance, translational regulation, quantitative systems biology, mass spectrometry, RNA decoding, protein synthesis and degradation, macrophage polarization, aging, neurodegenerative diseases
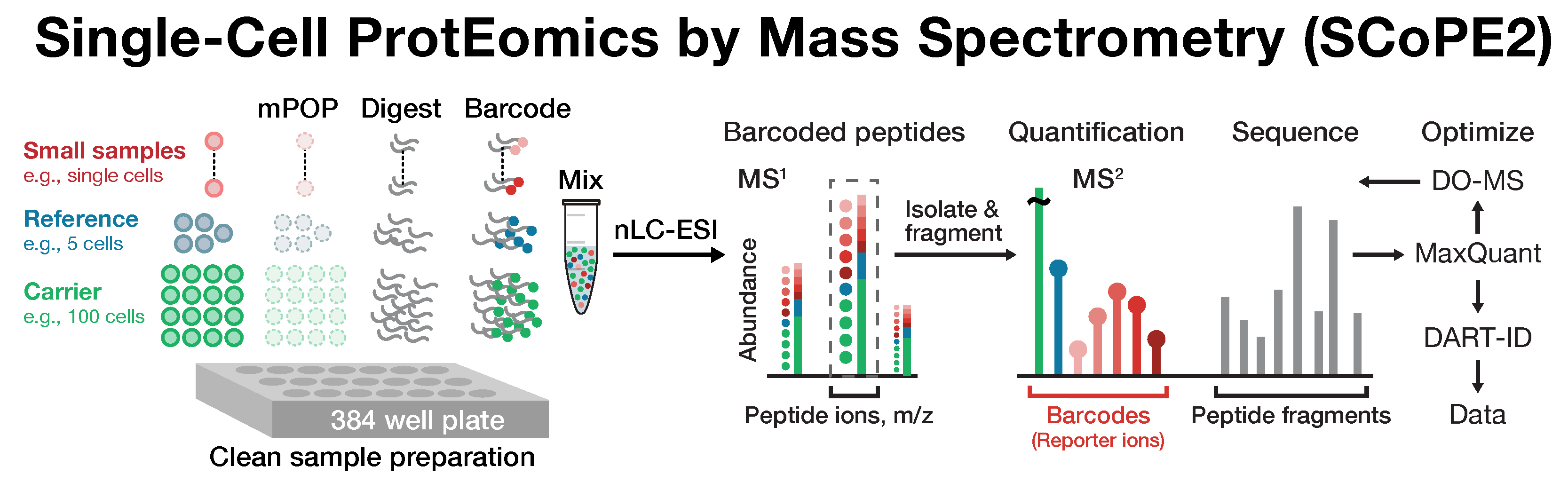
Single-cell proteomics by mass-spectrometry
Many biological processes stem from the coordinated interactions of molecularly and functionally diverse cells. However, this diversity is relatively unexplored at the proteome level because of the limitations of conventional affinity-based reagents for quantifying proteins in single cells. To alleviate these limitations, in 2017 our laboratory introduced Single Cell ProtEomics by Mass Spectrometry (SCoPE-MS). Since then, we developed more powerful and fully automated methods, including SCoPE2, pSCoPE, and plexDIA.
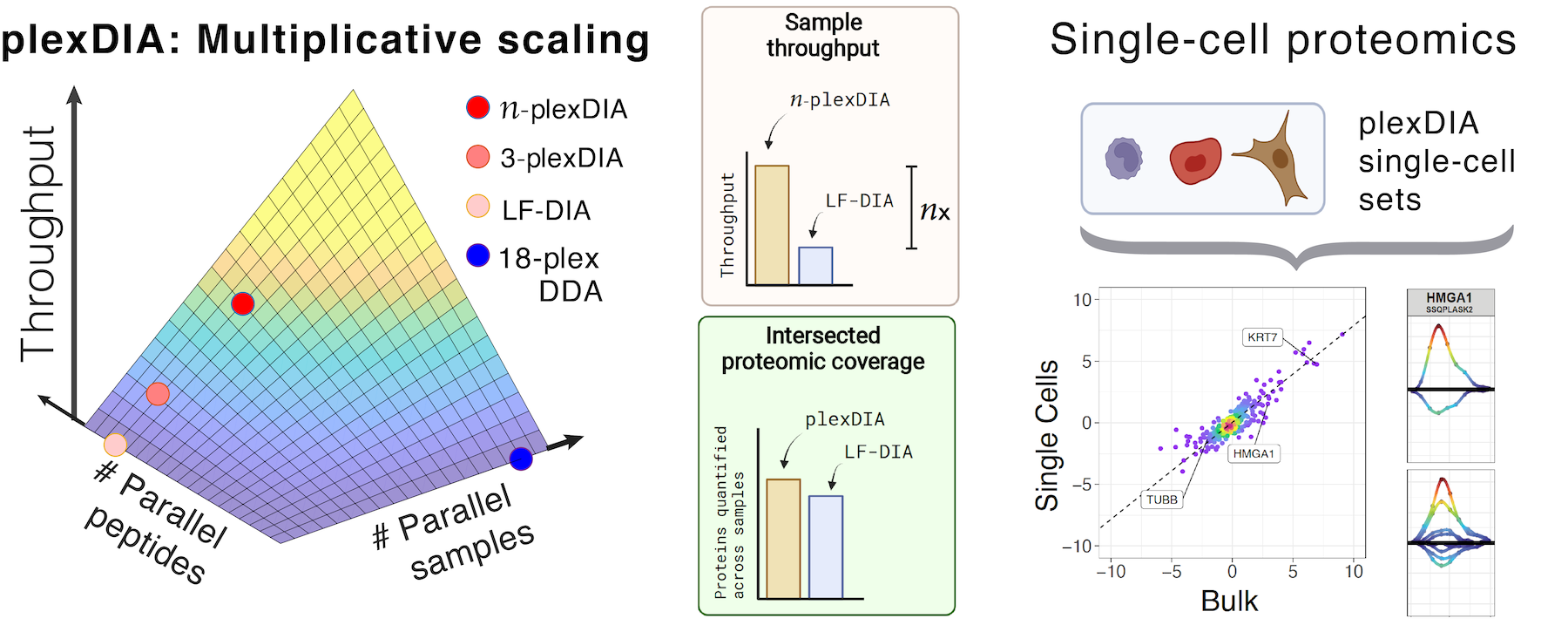
Taking advantage of strategies for advancing data acquisition and mass spectra identification, we developed methods that increase the sensitivity, data completeness and flexibility of single-cell protein analysis. These methods allow prioritization of thousands of proteins or parallelization in the mass domain by plexDIA, which supports parallel analysis of both single-cells and peptides. The multiplexing in the mass domain can be complemented by time domain multiplexing using timePlex, which enabled combinatorial increase in throughput as demonstrated by research at PTI. To facilitate accessibility, we introduced robust protocols and recommended best practices.
Dynamics of protein synthesis and degradation
Cellular protein concentrations are maintained through a balance of synthesis and clearance. Clearance occurs through both protein degradation and growth-dependent dilution. We have found that the impact of clearance in context-dependent and large, sometimes account for over 50 % of the measured variance in protein abundance. We study this contribution of protein degradation to shaping protein variation both across the proteome and across cell states.
Alternate RNA decoding
Amino acid substitutions may substantially alter protein stability and function, but the contribution of substitutions arising from alternate translation (deviations from the genetic code) is unknown. We have identified 60,024 high confidence substitutions corresponding to 8,801 unique sites in proteins from 1,990 genes. Some substitutions are shared across samples, while others exhibit strong tissue-type and cancer specificity. Surprisingly, products of alternate translation are more abundant than their canonical counterparts for hundreds of proteins, suggesting sense codon recoding. Recoded proteins include transcription factors, proteases, signaling proteins, and proteins associated with neurodegeneration. Mechanisms contributing to substitution abundance include protein stability, codon frequency, codon-anticodon mismatches, and RNA modifications. Some substitutions are shared across samples, while others exhibit strong tissue-type and cancer specificity. Surprisingly, products of alternate translation are more abundant than their canonical counterparts for hundreds of proteins, suggesting sense codon recoding. Recoded proteins include transcription factors, proteases, signaling proteins, and proteins associated with neurodegeneration. Mechanisms contributing to substitution abun-dance include protein stability, codon frequency, codon-anticodon mismatches, and RNA modifications. We characterize sequence motifs around alternatively translated amino acids and how substitution ratios vary across protein domains, tissue types and cancers. Both the sequence and the tissue-specificity of alternatively translated proteins are conserved between human and mouse. These results demonstrate the contribution of alternate translation to diversifying mammalian proteomes, and its association with protein stability, tissue-specific proteomes, and diseases.
Aging and neurodegenerative diseases
Molecular mechanisms that regulate protein synthesis and degradation are central to aging and neurodegenerative diseases because they determine the integrity, quality, and adaptability of the proteome over time. High fidelity protein synthesis is required for maintaining functional proteins and prevents the buildup of aberrant translation products, while efficient degradation pathways, such as the ubiquitin–proteasome system and the autophagy–lysosome axis, remove damaged, misfolded, or aggregated proteins. With aging, both synthesis fidelity and degradative capacity decline, leading to accumulation of toxic protein species, impaired cellular stress responses, and progressive loss of neuronal resilience. In neurodegenerative disorders like Alzheimer’s, Parkinson’s, and ALS, defects in these same pathways directly drive pathogenic protein aggregation, synaptic dysfunction, and neuronal death. Thus, maintaining balanced, high-fidelity protein turnover is essential for delaying aging phenotypes and preventing neurodegeneration.

Slavov Laboratory
We aim to understand the rules governing emergent systems-level behavior and to use these rules to rationally engineer biological systems. We make quantitative measurements, often at the single-cell level, to test different conceptual frameworks and discriminate among different classes of models.
Selected Research Projects
- Making Single-Cell Proteomics data FAIR
- Single-cell proteomic identification of novel markers of senescence
- Mapping the Transcriptome and Proteome of Human Testis in 3D
- Tracking proteome dynamics in single cells
- Alternate RNA decoding
- Ribosome-mediated translational regulation during stem cell differentiation
Research Centers and Institutes
Department Research Areas
Selected Publications
Key Research Articles
- Huffman RG, Leduc A, Wichmann C, … and Slavov N.✉
- Leduc A, Huffman RG, Cantlon J, Khan S, Slavov N.✉
- Exploring functional protein covariation across single cells using nPOP Genome Biol 23, 261 (2022) preprint | PDF | nPOP Web | Data | Tutorials | Highlight by GEN | Perspective | Conference Talk
- Derks J, Leduc A, Wallmann G, Huffman G, Willetts M, Khan S, … Slavov N.✉
- Increasing the throughput of sensitive proteomics by plexDIA, Nature Biotechnology (2022) preprint | OA | PDF | plexDIA web | SCP2022 Talk | Nature Research Briefing | Nature Blog | Highlight by Nature Methods
- Budnik B, Levy E, Harmange G, Slavov N.✉
- SCoPE-MS: mass-spectrometry of single mammalian cells quantifies proteome heterogeneity during cell differentiation Genome Biology 19:161 (2018) preprint | PDF | Data | CSHL Talk | C&EN Highlight
Recent Research Articles
- Slavov N.✉ (2026) Single-Cell Proteomic Technologies: Tools in the quest for principles, Annual Review Biophysics 55 preprint | PDF | Reviews
- Iwamoto-Stohl LK, Petelski AA, … Slavov N.✉, Zernicka-Goetz (2025) Fertilization triggers early proteomic symmetry breaking in mammalian embryos, Cell | preprint | PDF | Web | Video | Highlight
- Leduc A, Shipkovenska G, Xu Y, Franks A, Slavov N.✉ (2025) The regulation of protein abundance across single cells in a mammalian tissue, bioRxiv doi: 10.1101/2025.09.17.676955 | PDF | Data Web | Video
- Leduc A, Xu Y, Shipkovenska G, Dou Z, Slavov N✉ (2025) Limiting the impact of protein leakage in single-cell proteomics Nat Commun 16, 4169 doi: 10.1038/s41467-025-56736-7 | PDF | SI Web
- Leduc A, Slavov N.✉ (2025) Protein degradation and growth dependent dilution substantially shape mammalian proteomes, bioRxiv doi: 10.1101/2025.02.10.637566 | PDF | Github
- Tsour S, Machne R, Leduc A, Widmer S, Slavov N.✉ (2024) Alternate RNA decoding results in stable and abundant proteins in mammals bioRxiv doi: 10.1101/2024.08.26.609665 | PDF | Web | Video
- Leduc A, Khoury L., Cantlon J., Khan S, Slavov N.✉ (2024) Massively parallel sample preparation for multiplexed single-cell proteomics using nPOP Nature Protocols preprint | PDF | Web | Blog | Primer Video
- Galatidou S, Petelski A, Pujol A, Lattes K,… Slavov N✉, Barragan M (2024) Single-cell proteomics reveals decreased abundance of proteostasis and meiosis proteins in advanced maternal age oocytes, Molecular Human Reproduction doi: 10.1093/molehr/gaae023 | PDF | Data
- Rogers ZJ, Colombani T, Khan S, et al. (2024)
Controlling Pericellular Oxygen Tension in Cell Culture Reveals Distinct Breast Cancer Responses to Low Oxygen Tensions
Adv Sci. e2402557. doi: 10.1002/advs.202402557 | PDF - Khan S, Conover R, Asthagiri AR, Slavov N.✉ (2023) Dynamics of single-cell protein covariation during epithelial-mesenchymal transition J. Proteome Res. doi: 10.1021/acs.jproteome.4c00277 | PDF | Data Web
- Wallmann G., Leduc A., Slavov N.✉ (2023) Data-Driven Optimization of DIA Mass-Spectrometry by DO-MS J. Proteome Res. 22(10):3149-3158 OA | preprint | PDF | Code | Web
- Slavov N.✉ Single-cell proteomics: quantifying post-transcriptional regulation, Development 150, dev201492 (2023) PDF | Web Resources
- van den Berg P., Bérenger-Currias NM, Budnik B., Slavov N.✉, Semrau S.✉ (2023) Integration of a multi-omics stem cell differentiation dataset using a dynamical model PLoS Genet 19(5): e1010744 PDF | preprint | Data Web
- Huffman RG, Leduc A, Wichmann C, … and Slavov N.✉ (2023) Prioritized mass spectrometry increases the depth, sensitivity and data completeness of single-cell proteomics Nature Methods preprint | PDF | pSCoPE web | Data web | Talk | Nature Research Briefing (PDF)
- MacCoss MJ, Alfaro J, Faivre, D, Wu C, Wanunu M, Slavov N.✉ (2023) Sampling proteomes by emerging single-molecule and mass-spectrometry methods, Nature Methods 20, 339–346 preprint | PDF | SCP2022 Talk
- Gatto L, Aebersold R, Cox J, Demichev V, Derks J, …, Slavov N.✉ (2023) Initial recommendations for performing, benchmarking, and reporting single-cell proteomics experiments, Nature Methods preprint | PDF | Guidelines Web | Blog
- Derks J, Slavov N.✉ (2023) Strategies for increasing the depth and throughput of protein analysis by plexDIA, J. Proteome Res. preprint | PDF | Code | Web | YouTube Talk
- Leduc A, Huffman RG, Cantlon J, Khan S, Slavov N.✉ (2022) Exploring functional protein covariation across single cells using nPOP Genome Biol 23, 261 preprint | PDF | nPOP Web | Data | Tutorials | Highlight by GEN | Perspective | Conference Talk
- Derks J, Leduc A, Wallmann G, Huffman G, Willetts M, Khan S, Specht H, Ralser M, Demichev V, Slavov N.✉ (2022) Increasing the throughput of sensitive proteomics by plexDIA, Nature Biotechnology preprint | OA | PDF | plexDIA web | SCP2022 Talk | Nature Research Briefing | Nature Blog | Highlight by Nature Methods
- Slavov N.✉ (2022) Counting protein molecules for single-cell proteomics, Cell 185 (2) :232-234 PDF | OA
- Slavov N.✉ (2022) Learning from natural variation across the proteomes of single cells, PLoS Biology 20(1): e3001512 PDF | Website | Videos
- Slavov N.✉ (2021) Scaling up single-cell proteomics, Molecular & Cellular Proteomics PDF | Website | Recorded Workshop
- Petelski A, Emmott E, Leduc A, Huffman RG, Specht H, Perlman D, Slavov N.✉ (2021) Multiplexed single-cell proteomics using SCoPE2 Nature Protocols preprint | OA | PDF | Data web | Video Tutorials
- Slavov N.✉ (2021) Driving Single Cell Proteomics Forward with Innovation, J. of Proteome Res., doi: 10.1021/acs.jproteome.1c00639 | PDF
- Slavov N.✉ (2021) Increasing proteomics throughput, Nature Biotechnology PDF
- Slavov N.✉ et al. (2021) Voices of biotech research: Single-cell proteomics, Nature Biotechnology, 39, 281–286 PDF
- Specht H, Emmott E, Petelski A, Huffman RG, Perlman D, Serra M, Kharchenko P, Koller A. Slavov N.✉ (2021) Single-cell proteomic and transcriptomic analysis of macrophage heterogeneity using SCoPE2 Genome Biology preprint | PDF | SCP2019 talk | SCoPE2 Web
- Specht H, Slavov N.✉ (2020) Optimizing accuracy and depth of protein quantification in experiments using isobaric carriers, J. of Proteome Res. Preprint | PDF | WebSite | SCP2020 talk | Code
- Petelski AA, Slavov N.✉ (2020) Analyzing ribosome remodeling in health and disease, Proteomics preprint PDF
- Slavov N.✉ (2020) Single-cell protein analysis by mass-spectrometry, Current Opinion in Chemical Biology, 60, 1-9 PDF
- Slavov N.✉ (2020) Unpicking the proteome in single cells, Science, 367 (6477) :512–513
- Slavov N.✉ et al. (2019) Voices in methods development: Single-cell proteomics Nature Methods, PDF
- Specht H, Emmott E, Koller T, Slavov N.✉ (2019) High-throughput single-cell proteomics quantifies the emergence of macrophage heterogeneity
bioRxiv DOI: 10.1101/665307 PDF | Data | SCP2019 talk | SCoPE2 Web - Chen A, Franks A, Slavov N.✉ (2019) DART-ID increases single-cell proteome coverage, PLoS Computational Biology, DOI: 10.1371/journal.pcbi.1007082 | PDF | RAW Data @ MassIVE | GitHub | DART-ID web
- Huffman RG, Specht H, Chen AT, Slavov N. (2019) DO-MS: Data-Driven Optimization of Mass Spectrometry Methods, J. of Proteome Res., DOI: 10.1021/acs.jproteome.9b00039 | DO-MS web | DO-MS @ GitHub
- Budnik B, Levy E, Harmange G, Slavov N.✉ (2018)
SCoPE-MS: mass-spectrometry of single mammalian cells quantifies proteome heterogeneity during cell differentiation
Genome Biology 19:161 preprint | PDF | Data | CSHL Talk | C&EN Highlight - Emmott EP, Jovanovic M, Slavov N.✉ (2018) Ribosome stoichiometry: from form to function, Trends in Biomedical Sciences, DOI: 10.1016/j.tibs.2018.10.009 PDF
- Malioutov D., Chen T., Jaffe J., Airoldi E., Budnik B., Slavov N. ✉ (2018) Quantifying homologous proteins and proteoforms, Molecular & Cellular Proteomics, DOI: 10.1101/168765 | PDF
- Specht H, Harmange G, Perlman DH, Emmott E, Niziolek Z, Budnik B, Slavov N.✉Automated sample preparation for high-throughput single-cell proteomics, bioRxiv, DOI: 10.1101/399774 PDF | RAW Data @ MassIVE | SCP2018 Talk
- Specht H. & Slavov N. (2018) Transformative opportunities for single cell proteomics J. of Proteome Res., 17 (8), 2565 – 2571
- Berg P., Budnik B., Slavov N., Semrau S. (2017) Dynamic post-transcriptional regulation during embryonic stem cell differentiation, bioRxiv, DOI: 10.1101/123497
- Franks A., Airoldi E.M., Slavov N. (2017) Post-transcriptional regulation across human tissues, PLoS Computational Biology, 13(5): e1005535, DOI: 10.1101/020206 | Highlight by The Scientist
- Slavov N. (2015) Making the most of peer review, eLife, 4:e12708
- Slavov N., Semrau S., Airoldi E., Budnik B., van Oudenaarden A., (2015) Differential Stoichiometry among Core Ribosomal Proteins, Cell Reports, 13(5), 865-873. (TiBS Spotlight)
- Malioutov D., Slavov N. (2014) Convex Total Least Squares, Journal of Machine Learning Research, W&CP, 32(1), 109-117.
- Slavov N., Budnik B., Schwab D., Airoldi E., van Oudenaarden A. (2014) Constant Growth Rate Can Be Supported by Decreasing Energy Flux and Increasing Aerobic Glycolysis, Cell Reports 7(3), 705-714.
- Slavov N., Carey, J., Linse, S. (2013) Calmodulin transduces Ca+2 oscillations into differential regulation of its target proteins, ACS Chemical Neuroscience, 4, 601-612.
- Slavov N., Botstein D. (2013) Decoupling Nutrient Signaling from Growth Rate Causes Aerobic Glycolysis and Deregulation of Cell Size and Gene Expression, Molecular Biology of the Cell, 24(2), 157-168,
- Slavov N., van Oudenaarden A. (2012) How to Regulate a Gene: To Repress or to Activate?, Molecular Cell, 46(5), 551-552.
- Slavov N., Airoldi E.M., van Oudenaarden A., Botstein D. (2012) A Conserved Cell Growth Cycle Can Account for the Environmental Stress Responses of Divergent Eukaryotes, Mol. Biol. Cell, vol. 23, no. 10
- Slavov N., Macinskas J., Caudy A., Botstein D. (2011) Metabolic Cycling without Cell Division Cycling in Respiring Yeast, PNAS, vol. 108, 19090-19095
- Slavov N., Botstein D. (2011) Coupling among Growth Rate Response, Metabolic Cycle and Cell Division Cycle in Yeast, Mol. Biol. Cell, vol. 22
- Slavov N. (2010) Inference of Sparse Networks with Unobserved Variables. Application to Gene Regulatory Networks, JMLR, W&CP vol. 9
- Slavov, N., Dawson, K. (2009) Correlation Signature of the Macroscopic States of the Gene Regulatory Network in Cancer, PNAS, vol. 106, no. 11 PDF

Apr 29, 2026
Slavov Selected for ASMS Biemann Medal
BioE Professor Nikolai Slavov was selected to receive the 2026 Biemann Medal from the American Society for Mass Spectrometry (ASMS), which is awarded to an individual early in their career in recognition of significant achievement in basic or applied mass spectrometry and is among the most prestigious awards in that field.

Jan 29, 2026
Faculty and Staff Awards 2026
Faculty and staff were recognized at the 28th Annual College of Engineering Faculty and Staff Awards for their exceptional service and dedication in support of students, the COE community, and the university during the 2025-2026 academic year.

Dec 04, 2025
New Discovery in Human Cells
BioE Professor Nikolai Slavov and his team at the Single-Cell Proteomics Center made a discovery in early human cell differences. The findings of this discovery could have a huge impact on regenerative medicine.

Oct 24, 2025
When Variant Proteins Aren’t Actually the Variant Ones
The bioRxiv preprint of BioE Professor Nikolai Slavov’s research group on “Alternate RNA decoding results in stable and abundant proteins in mammals” was featured in the Science article “When Variant Proteins Aren’t Actually the Variant Ones.”
Sep 23, 2025
2025 Stanford University Annual Assessment of Author Citations
The following COE professors are among the top scientists worldwide selected by Stanford University representing the top 2% of the most-cited scientists with single-year impact in various disciplines. The selection is based on the top 100,000 by c-score (with and without self-citations) or a percentile rank of 2% or above.

Aug 08, 2025
Decoding the Proteins Inside Every Cell
BioE Professor Nikolai Slavov, in collaboration with EMBL-EBI, Cambridge, UK, was awarded a £2,000,000 NSF grant for “Making Single-Cell Proteomics data FAIR.”

May 27, 2025
Addressing Protein Leakage in Cells To Improve the Accuracy of Single-Cell Proteomic Data
Research conducted by BioE Professor Nikolai Slavov’s group was featured in the AZO Life Sciences article titled “Addressing Protein Leakage in Cells To Improve the Accuracy of Single-Cell Proteomic Data.”
Jan 31, 2025
2024 Stanford University Annual Assessment of Author Citations
The following COE professors are among the top scientists worldwide selected by Stanford University representing the top 2% of the most-cited scientists with single-year impact in various disciplines. The selection is based on the top 100,000 by c-score (with and without self-citations) or a percentile rank of 2% or above. The list below includes those who published a paper in 2024 or later.

Feb 26, 2024
Northeastern Among Top 100 Universities for Utility Patents
Northeastern University, recognized for its excellence in innovation and entrepreneurship, has once again been named one of the top 100 universities worldwide for securing utility patents by the National Academy of Inventors.

Jan 22, 2024
Addressing Risk Factors and Potential Treatments of Alzheimer’s and Dementia
BioE Associate Professor Nikolai Slavov and Bouve Associate Professor Becky Briesacher provide insight into the differences between Alzheimer’s and dementia.